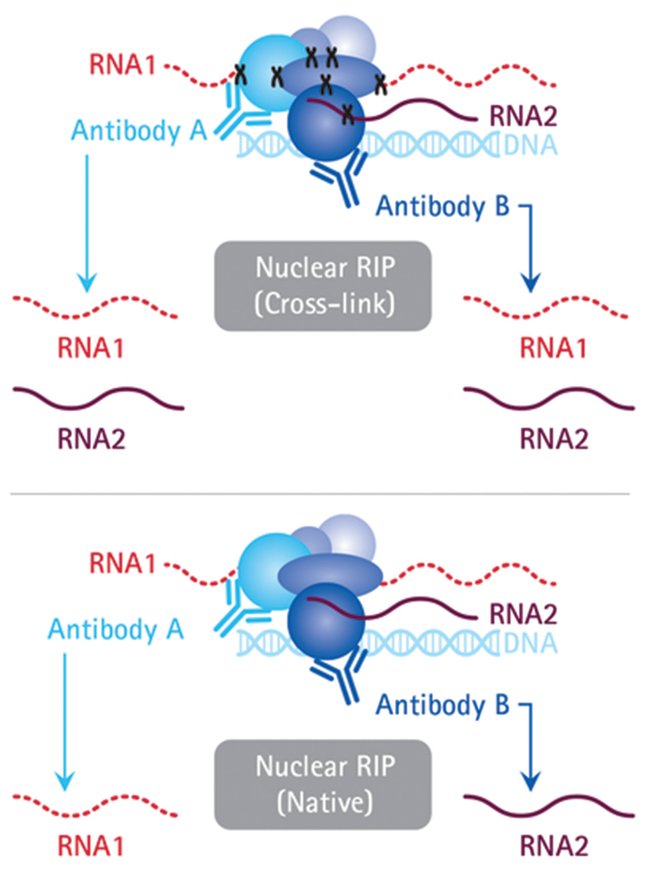
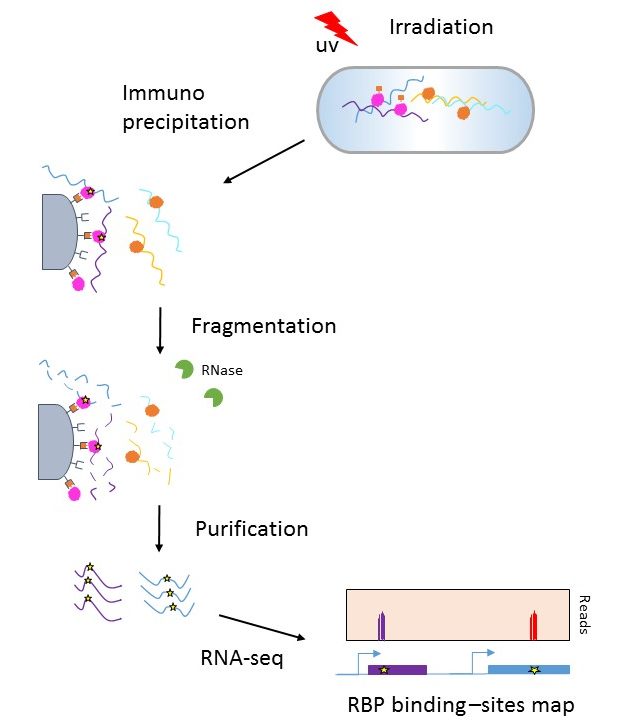
Enhanced immunoprecipitation techniques for the identification of RNA-binding protein partners: IGF2BP1 interactions in mammary epithelial cells - Journal of Biological Chemistry

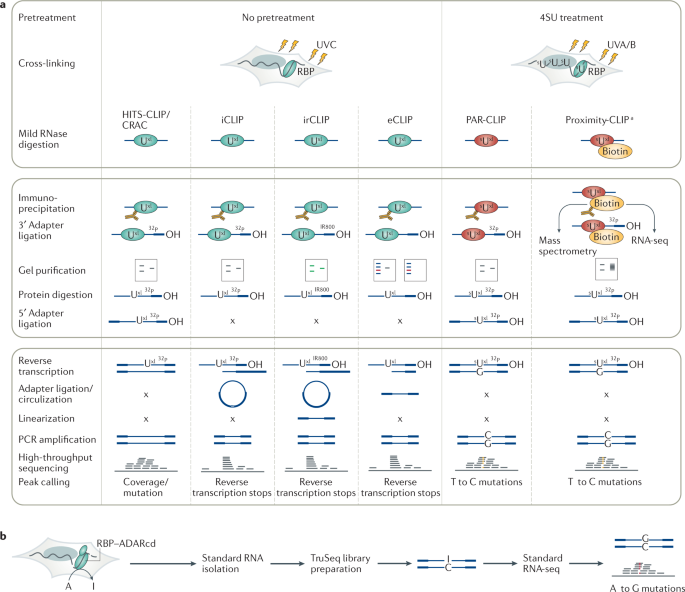
Global Approaches in Studying RNA-Binding Protein Interaction Networks: Trends in Biochemical Sciences

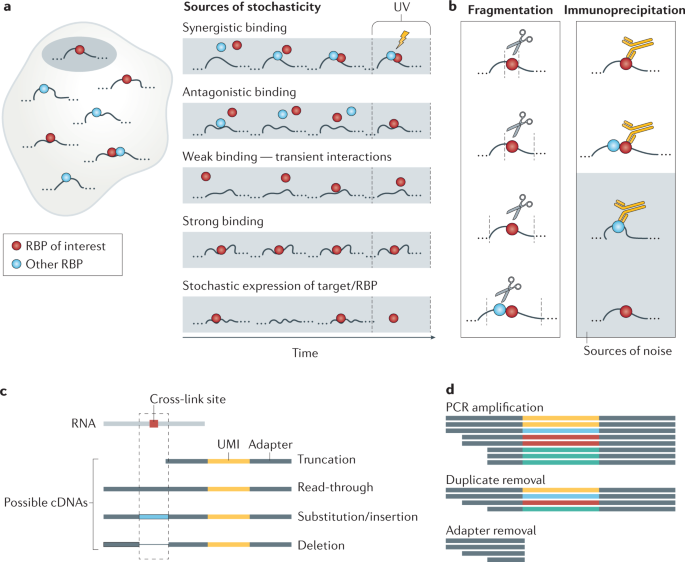
Experimental and Computational Considerations in the Study of RNA-Binding Protein-RNA Interactions. - Abstract - Europe PMC

Computational approaches for the analysis of RNA–protein interactions: A primer for biologists - Journal of Biological Chemistry
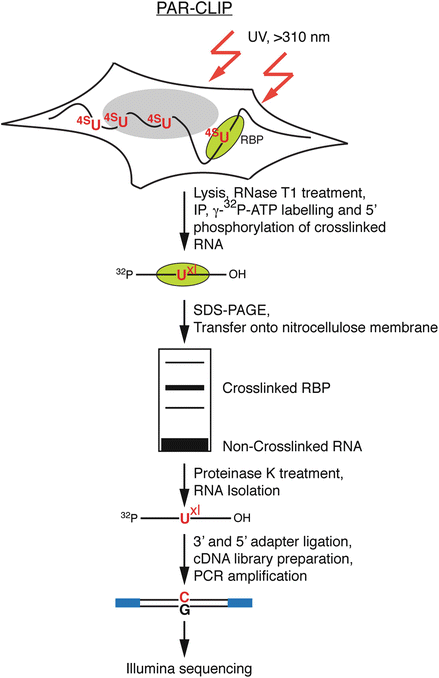
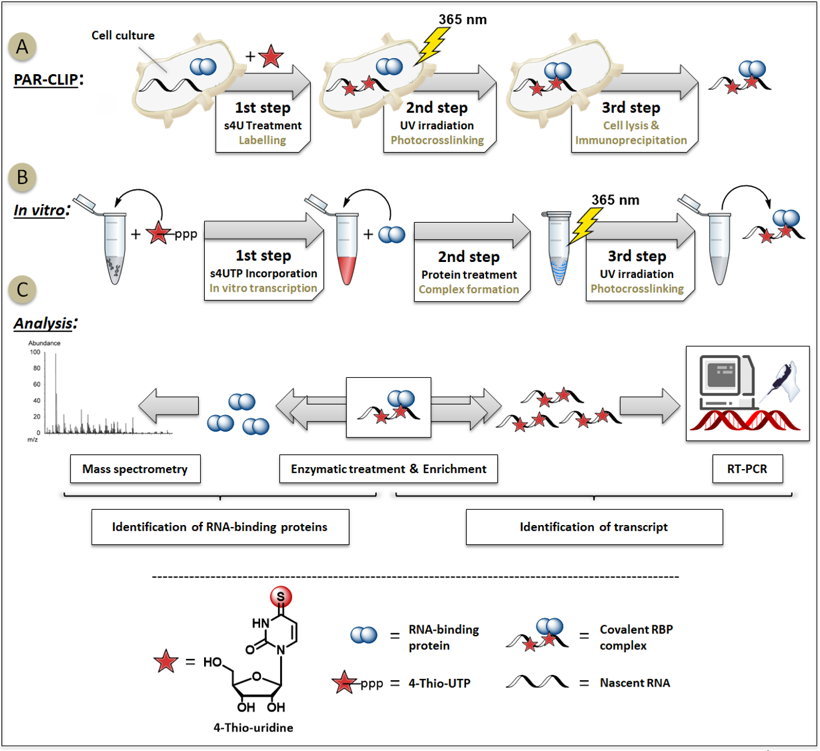
PAR-CLIP: A Method for Transcriptome-Wide Identification of RNA Binding Protein Interaction Sites | SpringerLink
PRAS: Predicting functional targets of RNA binding proteins based on CLIP-seq peaks | PLOS Computational Biology
![PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/85f2546f2ebc745edebbaf20459e20a394664b72/5-Figure4-1.png)
PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar

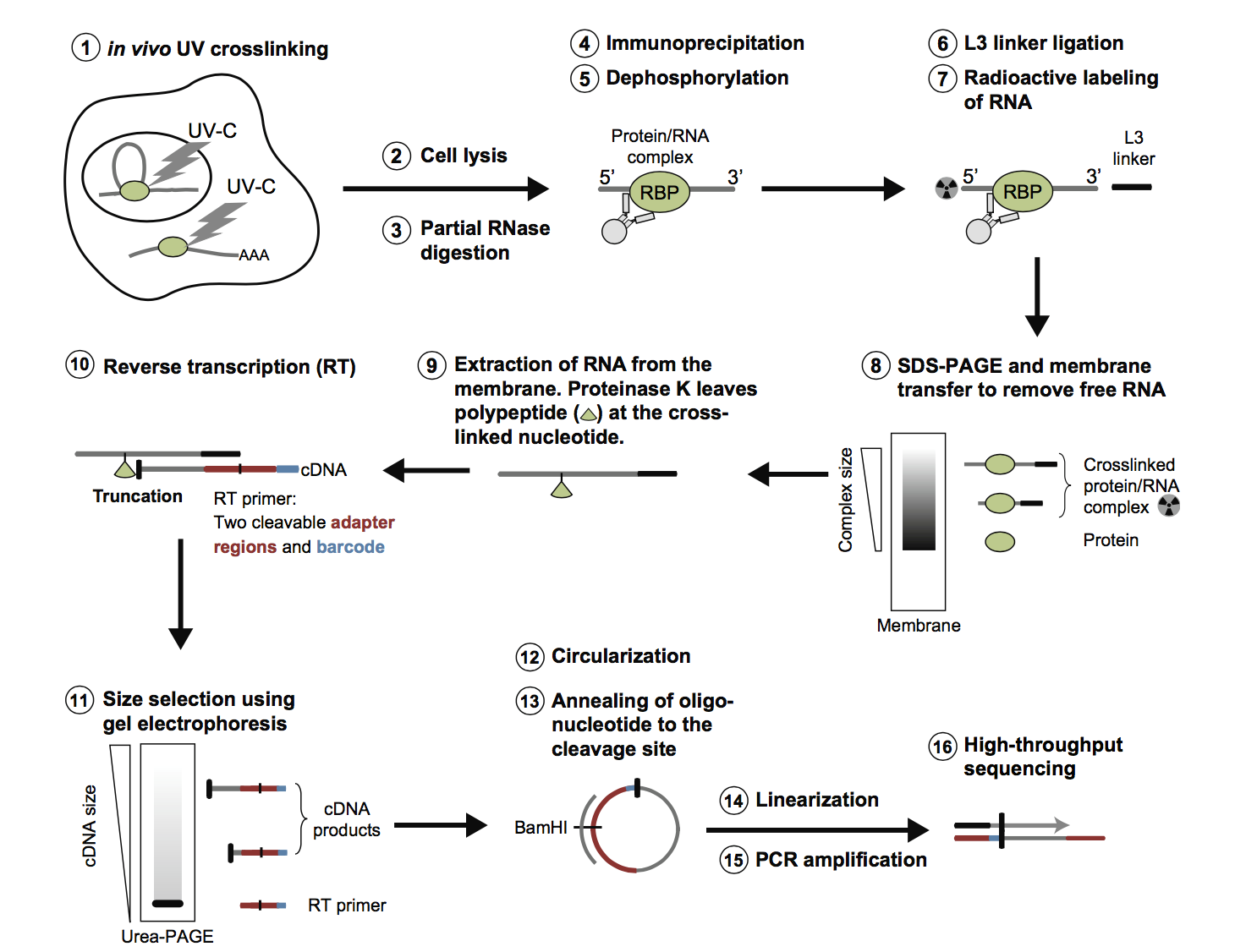
PAR-CLIP and Streamlined Small RNA cDNA Library Preparation Protocol for the Identification of RNA Binding Protein Target Sites | RNA-Seq Blog

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell
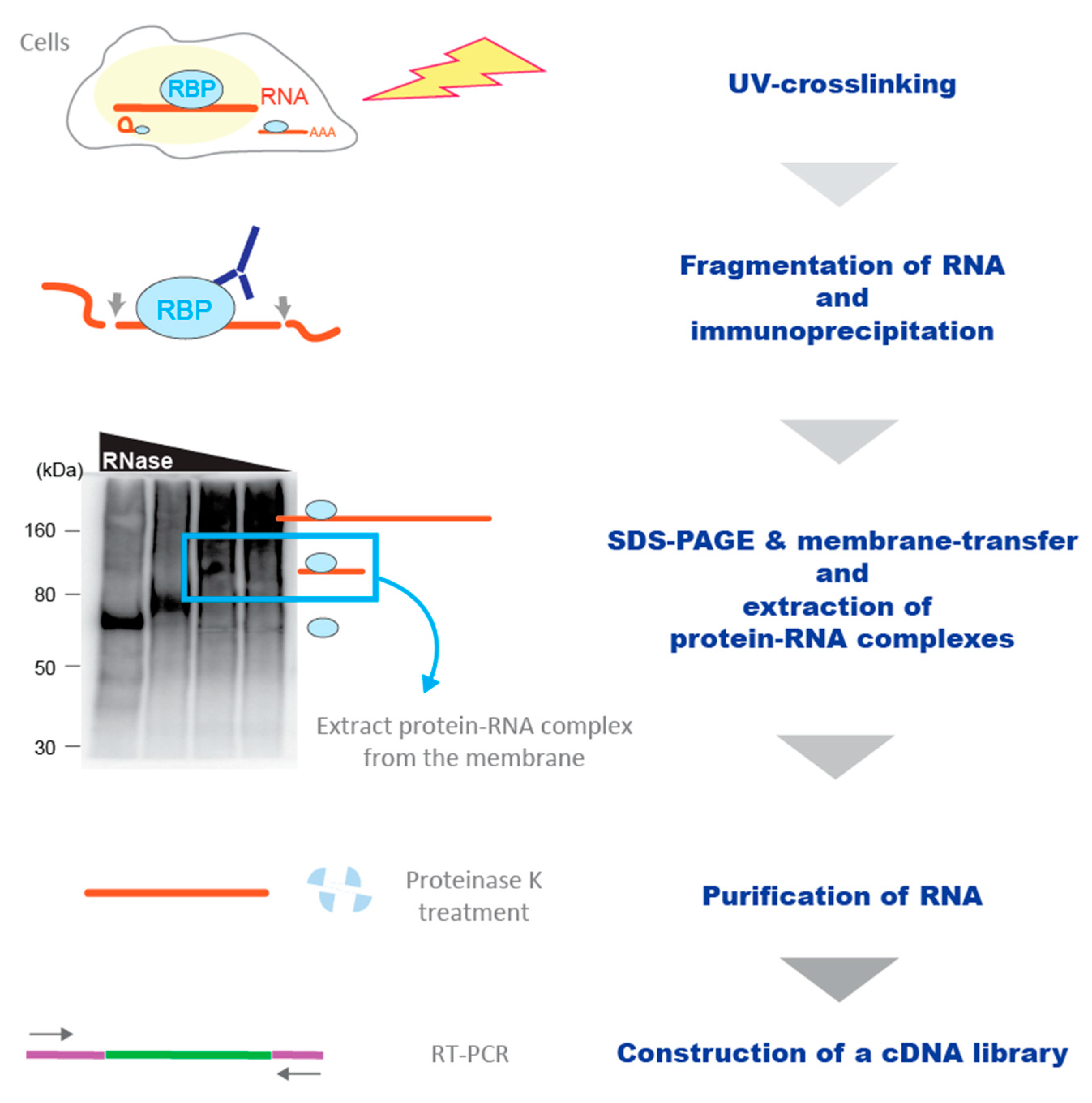
IJMS | Free Full-Text | Rapidly Growing Protein-Centric Technologies to Extensively Identify Protein–RNA Interactions: Application to the Analysis of Co-Transcriptional RNA Processing | HTML

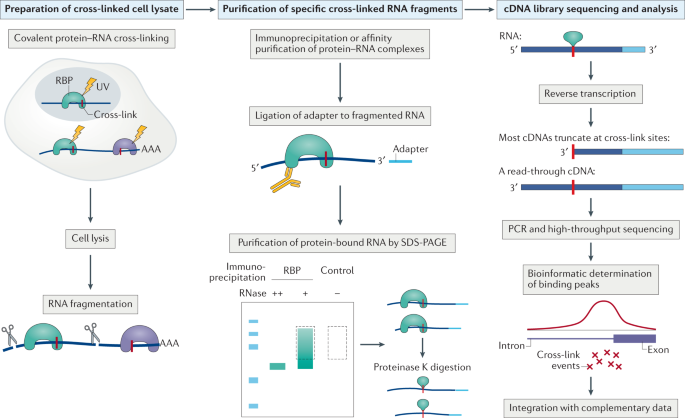
Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram




-2.jpg)